Nevertheless, SBT is unable to differentiate Lp strains with locally prevalent STs, thus creating uncertainty around the interpretation of isolate genetic relationships. pneumophila DNA molecular typing over the past decade, allowing for universal exchange of sequence type (ST) information.

While both are currently in use, SBT became the gold standard for L. Thus, molecular comparison of clinical and environmental isolates is helpful for source attribution during LD outbreaks at least two laboratory techniques, pulsed-field gel electrophoresis (PFGE) and sequence-based typing (SBT), have been widely used for this purpose. pneumophila (Lp), and up to 79% of Lp infections are attributable to serogroup 1 (Lp1). Approximately 80% of laboratory diagnosed LD in the United States (US) is due to a single species, L. Legionella is a globally important cause of severe and sometimes fatal bacterial pneumonia known as Legionnaires’ disease (LD). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist. No additional external funding was received for this study. Raw Illumina sequencing reads were assigned the SRA accession SRP127407 (Sequence Read Archive, ) and individual isolate SRA sequence accession IDs are listed in S1 Table.įunding: This study was supported by funding made available through the CDC Office of Advanced Molecular Detection. The work is made available under the Creative Commons CC0 public domain dedication.ĭata Availability: Sequencing data derived from this study have been deposited with links to BioProject accession number PRJNA423272 in the NCBI BioProject Database ( ).
PBP3 GENE AND LEGIONELLA FREE
This is an open access article, free of all copyright, and may be freely reproduced, distributed, transmitted, modified, built upon, or otherwise used by anyone for any lawful purpose.

Received: JAccepted: OctoPublished: October 18, 2018 Ashley Robinson, University of Mississippi Medical Center, UNITED STATES These findings provide new insight into the ST1 population structure and establish a foundation for interpreting genetic relationships among ST1 strains these data may also inform future analyses for improved outbreak investigations.Ĭitation: Mercante JW, Caravas JA, Ishaq MK, Kozak-Muiznieks NA, Raphael BH, Winchell JM (2018) Genomic heterogeneity differentiates clinical and environmental subgroups of Legionella pneumophila sequence type 1. Up to ~10% of US ST1 genetic variation could be explained by geographic origin, but considerable genetic conservation existed among strains isolated from geographically distant states and from different decades.

The accessory pangenome of environmental isolates was also ~30–60% larger than other subgroups and was enriched for transposition and conjugative transfer-associated elements. The ST1 population was highly conserved at the nucleotide level 98% of core nucleotide positions were invariant and environmental isolates unassociated with human disease (n = 99) contained ~65% more nucleotide diversity compared to clinical-sporadic (n = 139) or outbreak-associated (n = 28) ST1 subgroups. In order to characterize the ST1 population, we sequenced 289 outbreak and non-outbreak associated clinical and environmental ST1 and ST1-variant Lp strains from the US and, together with international isolate sequences, explored their genetic and geographic diversity.
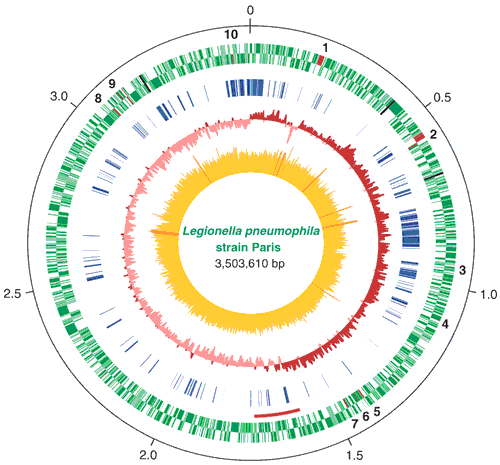
In the United States (US), ST1 is the most prevalent clinical and environmental Lp sequence type. pneumophila (Lp) sequence types (ST), which complicates laboratory investigations and environmental source attribution. In some cases, current genetic subtyping methods cannot resolve LD outbreaks caused by common, potentially endemic L. are the cause of a severe bacterial pneumonia known as Legionnaires’ disease (LD).
